Spectral fitting example (Crab)
This notebook fits the spectrum of a Crab simulated using MEGAlib and combined with background.
3ML is a high-level interface that allows multiple datasets from different instruments to be used coherently to fit the parameters of source model. A source model typically consists of a list of sources with parametrized spectral shapes, sky locations and, for extended sources, shape. Polarization is also possible. A “coherent” analysis, in this context, means that the source model parameters are fitted using all available datasets simultanously, rather than performing individual fits and finding a well-suited common model a posteriori.
In order for a dataset to be included in 3ML, each instrument needs to provide a “plugin”. Each plugin is responsible for reading the data, convolving the source model (provided by 3ML) with the instrument response, and returning a likelihood. In our case, we’ll compute a binned Poisson likelihood:
where \(d_i\) are the counts on each bin and \(\lambda_i\) are the expected counts given a source model with parameters \(\mathbf{x}\).
In this example, we will fit a single point source with a known location. We’ll assume the background is known and fixed up to a scaling factor. Finally, we will fit a Band function:
where \(K\) (normalization), \(\alpha\) & \(\beta\) (spectral indeces), and \(x_p\) (peak energy) are the free parameters, while \(E_{piv}\) is the pivot energy which is fixed (and arbitrary).
Considering these assumptions:
where \(B*b_i\) are the estimated counts due to background in each bin with \(B\) the amplitude and \(b_i\) the shape of the background, and \(s_i\) are the corresponding expected counts from the source, the goal is then to find the values of \(\mathbf{x} = [K, \alpha, \beta, x_p]\) and \(B\) that maximize \(\mathcal{L}\). These are the best estimations of the parameters.
The final module needs to also fit the time-dependent background, handle multiple point-like and extended sources, as well as all the spectral models supported by 3ML. Eventually, it will also fit the polarization angle. However, this simple example already contains all the necessary pieces to do a fit.
[1]:
from cosipy import BinnedData
from cosipy.spacecraftfile import SpacecraftHistory
from cosipy.response.FullDetectorResponse import FullDetectorResponse
from cosipy.util import fetch_wasabi_file
from cosipy.statistics import PoissonLikelihood
from cosipy.background_estimation import FreeNormBinnedBackground
from cosipy.interfaces import ThreeMLPluginInterface
from cosipy.response import BinnedThreeMLModelFolding, BinnedInstrumentResponse, BinnedThreeMLPointSourceResponse
from cosipy.data_io import EmCDSBinnedData
import sys
import astropy.units as u
import numpy as np
import matplotlib.pyplot as plt
from threeML import Band, PointSource, Model, JointLikelihood, DataList
from astromodels import Parameter, Powerlaw
from pathlib import Path
%matplotlib inline
15:27:55 WARNING The naima package is not available. Models that depend on it will not be functions.py:43 available
WARNING The GSL library or the pygsl wrapper cannot be loaded. Models that depend on it functions.py:65 will not be available.
WARNING The ebltable package is not available. Models that depend on it will not be absorption.py:33 available
15:27:56 INFO Starting 3ML! __init__.py:44
WARNING WARNINGs here are NOT errors __init__.py:45
WARNING but are inform you about optional packages that can be installed __init__.py:46
WARNING to disable these messages, turn off start_warning in your config file __init__.py:47
WARNING ROOT minimizer not available minimization.py:1208
WARNING Multinest minimizer not available minimization.py:1218
WARNING PyGMO is not available minimization.py:1228
WARNING The cthreeML package is not installed. You will not be able to use plugins which __init__.py:95 require the C/C++ interface (currently HAWC)
WARNING Could not import plugin HAWCLike.py. Do you have the relative instrument __init__.py:136 software installed and configured?
WARNING Could not import plugin FermiLATLike.py. Do you have the relative instrument __init__.py:136 software installed and configured?
WARNING No fermitools installed lat_transient_builder.py:44
WARNING Env. variable OMP_NUM_THREADS is not set. Please set it to 1 for optimal __init__.py:345 performances in 3ML
WARNING Env. variable MKL_NUM_THREADS is not set. Please set it to 1 for optimal __init__.py:345 performances in 3ML
WARNING Env. variable NUMEXPR_NUM_THREADS is not set. Please set it to 1 for optimal __init__.py:345 performances in 3ML
Download and read in binned data
Define the path to the directory containing the data, detector response, orientation file, and yaml files if they have already been downloaded, or the directory to download the files into
[2]:
data_path = Path("") # /path/to/files. Current dir by default
Download the orientation file (684.38 MB)
[3]:
fetch_wasabi_file('COSI-SMEX/DC2/Data/Orientation/20280301_3_month_with_orbital_info.ori', output=str(data_path / '20280301_3_month_with_orbital_info.ori'), checksum = '416fcc296fc37a056a069378a2d30cb2')
A file named 20280301_3_month_with_orbital_info.ori already exists with the specified checksum (416fcc296fc37a056a069378a2d30cb2). Skipping.
Read in the spacecraft orientation file
[4]:
sc_orientation = SpacecraftHistory.open(data_path / "20280301_3_month_with_orbital_info.ori")
Download signal and background data. FreeNormBinnedBackground supports multiple components, the normalization of all of them would be fitted simultanously. To make this example run fast, the default is to combined all of them into a single background model.
[5]:
single_bkg_fit = True
crab_data_path = data_path / "crab_standard_3months_binned_data_filtered_with_SAAcut.fits.gz.hdf5"
fetch_wasabi_file('COSI-SMEX/cosipy_tutorials/crab_spectral_fit_galactic_frame/crab_standard_3months_binned_data_filtered_with_SAAcut.fits.gz.hdf5',
output=crab_data_path, checksum = '405862396dea2be79d7892d6d5bb50d8')
bkg_components = {"PrimaryProtons":{'filename':'PrimaryProtons_WithDetCstbinned_data_filtered_with_SAAcut.hdf5', 'checksum':'7597f04210e59340a0888c66fc5cbc63'},
"PrimaryAlphas": {'filename': 'PrimaryAlphas_WithDetCstbinned_data_filtered_with_SAAcut.hdf5', 'checksum': '76a68da730622851b8e1c749248c3b40'},
"AlbedoPhotons": {'filename': 'AlbedoPhotons_WithDetCstbinned_data_filtered_with_SAAcut.hdf5', 'checksum': '76c58361d2c9b43b66ef2e41c18939c4'},
"AlbedoNeutrons": {'filename': 'AlbedoNeutrons_WithDetCstbinned_data_filtered_with_SAAcut.hdf5', 'checksum': '8f3cb418c637b839665a4fcbd000d2eb'},
"CosmicPhotons": {'filename': 'CosmicPhotons_3months_binned_data_filtered_with_SAAcut.hdf5', 'checksum': '93c4619b383572d318328e6380e35a70'},
"CosmicDiffuse": {'filename': 'GalTotal_SA100_F98_3months_binned_data_filtered_with_SAAcut.hdf5', 'checksum': 'd0415d4d04b040af47f23f5d08cb7d64'},
"SecondaryPositrons": {'filename': 'SecondaryPositrons_3months_binned_data_filtered_with_SAAcut.hdf5', 'checksum': '5fec2212dcdbb4c43c3ac02f02524f68'},
"SecondaryProtons": {'filename': 'SecondaryProtons_WithDetCstbinned_data_filtered_with_SAAcut.fits.gz.hdf5', 'checksum': '78aefa46707c98563294a898a62845c1'},
"SAAprotons": {'filename': 'SAA_3months_unbinned_data_filtered_with_SAAcut_statreduced_akaHEPD01result.hdf5', 'checksum': 'fc69fbbfd94cd595f57a8b11fc721169'},
}
# Download the binned background data
for bkg in bkg_components.values():
wasabi_path = 'COSI-SMEX/cosipy_tutorials/crab_spectral_fit_galactic_frame/'+bkg['filename']
fetch_wasabi_file(wasabi_path, output=data_path/bkg['filename'], checksum = bkg['checksum'])
A file named crab_standard_3months_binned_data_filtered_with_SAAcut.fits.gz.hdf5 already exists with the specified checksum (405862396dea2be79d7892d6d5bb50d8). Skipping.
A file named PrimaryProtons_WithDetCstbinned_data_filtered_with_SAAcut.hdf5 already exists with the specified checksum (7597f04210e59340a0888c66fc5cbc63). Skipping.
A file named PrimaryAlphas_WithDetCstbinned_data_filtered_with_SAAcut.hdf5 already exists with the specified checksum (76a68da730622851b8e1c749248c3b40). Skipping.
A file named AlbedoPhotons_WithDetCstbinned_data_filtered_with_SAAcut.hdf5 already exists with the specified checksum (76c58361d2c9b43b66ef2e41c18939c4). Skipping.
A file named AlbedoNeutrons_WithDetCstbinned_data_filtered_with_SAAcut.hdf5 already exists with the specified checksum (8f3cb418c637b839665a4fcbd000d2eb). Skipping.
A file named CosmicPhotons_3months_binned_data_filtered_with_SAAcut.hdf5 already exists with the specified checksum (93c4619b383572d318328e6380e35a70). Skipping.
A file named GalTotal_SA100_F98_3months_binned_data_filtered_with_SAAcut.hdf5 already exists with the specified checksum (d0415d4d04b040af47f23f5d08cb7d64). Skipping.
A file named SecondaryPositrons_3months_binned_data_filtered_with_SAAcut.hdf5 already exists with the specified checksum (5fec2212dcdbb4c43c3ac02f02524f68). Skipping.
A file named SecondaryProtons_WithDetCstbinned_data_filtered_with_SAAcut.fits.gz.hdf5 already exists with the specified checksum (78aefa46707c98563294a898a62845c1). Skipping.
A file named SAA_3months_unbinned_data_filtered_with_SAAcut_statreduced_akaHEPD01result.hdf5 already exists with the specified checksum (fc69fbbfd94cd595f57a8b11fc721169). Skipping.
Load binned .hdf5 files. Create BinnedData objects for the Crab only, Crab+background, and background only. The Crab only simulation is not used for the spectral fit, but can be used to compare the fitted spectrum to the source simulation
[6]:
crab = BinnedData(data_path / "crab.yaml")
crab.load_binned_data_from_hdf5(binned_data=crab_data_path)
for bkg in bkg_components.values():
binned_data = BinnedData(data_path / "background.yaml")
binned_data.load_binned_data_from_hdf5(binned_data=data_path/bkg['filename'])
bkg['dist'] = binned_data.binned_data.project('Em', 'Phi', 'PsiChi')
total_bkg = None
for bkg in bkg_components.values():
if total_bkg is None:
total_bkg = bkg['dist']
else:
total_bkg = total_bkg + bkg['dist'] # Issues with in-place operations for sparse contents
if single_bkg_fit:
bkg_dist = {"total_bkg":total_bkg}
else:
bkg_dist = {l: b['dist'] for l, b in bkg_components.items()}
# Workaround to avoid inf values. Out bkg should be smooth, but currently it's not.
# Reproduces results before refactoring. It's not _exactly_ the same, since this fudge value was 1e-12, and
# it was added to the expectation, not the normalized bkg
for bckfile in bkg_dist.keys() :
bkg_dist[bckfile] += sys.float_info.min
Fetch and open the response
[7]:
dr_path = str(data_path / "ResponseContinuum.o3.e100_10000.b10log.s10396905069491.m2284.filtered.nonsparse.binnedimaging.imagingresponse.h5") # path to detector response
fetch_wasabi_file('COSI-SMEX/develop/Data/Responses/ResponseContinuum.o3.e100_10000.b10log.s10396905069491.m2284.filtered.nonsparse.binnedimaging.imagingresponse.h5',
output=str(dr_path), checksum = '7121f094be50e7bfe9b31e53015b0e85')
dr = FullDetectorResponse.open(dr_path)
A file named ResponseContinuum.o3.e100_10000.b10log.s10396905069491.m2284.filtered.nonsparse.binnedimaging.imagingresponse.h5 already exists with the specified checksum (7121f094be50e7bfe9b31e53015b0e85). Skipping.
Perform spectral fit
Set background parameter, which is used to fit the amplitude of the background, and instantiate the COSI 3ML plugin
[8]:
# Wrap the raw BinnedData objects into the appropiate data interface.
data = EmCDSBinnedData(crab.binned_data.project('Em', 'Phi', 'PsiChi') + total_bkg)
# Use the background model to initialize a background expectation interface.
# For this particular background interface implementation, only the normalization values are free.
bkg = FreeNormBinnedBackground(bkg_dist,
sc_history=sc_orientation,
copy = False)
# Wrape the raw response with BinnedInstrumentResponseInterface implementation
instrument_response = BinnedInstrumentResponse(dr, data)
# Initialize the 3ML Point Source response
# Note: Currently we're using the same NuLambda, Ei and Pol axes as the underlying FullDetectorResponse,
# matching the behavior of v0.3. This is all the current BinnedInstrumentResponse can do.
# In principle, this can be decoupled, and a BinnedInstrumentResponseInterface implementation
# can provide the response for an arbitrary directions, Ei and Pol values.
psr = BinnedThreeMLPointSourceResponse(data = data,
instrument_response = instrument_response,
sc_history=sc_orientation,
energy_axis = dr.axes['Ei'],
polarization_axis = dr.axes['Pol'] if 'Pol' in dr.axes.labels else None,
nside = 2*data.axes['PsiChi'].nside)
# Pass the 3ML Point Source response to interface implementation that will peform the
# folding with the spectrum
response = BinnedThreeMLModelFolding(data = data, point_source_response = psr)
# Likelihood to use
like_fun = PoissonLikelihood(data, response, bkg)
# Init 3ML plugin
cosi = ThreeMLPluginInterface('cosi',
like_fun,
response,
bkg)
# Init background parameters, consider as "nuisance parameters"
for bkg_label in bkg_dist.keys():
cosi.bkg_parameter[bkg_label] = Parameter(bkg_label, # background parameter
1, # initial value of parameter
min_value=0, # minimum value of parameter
max_value= 100 if single_bkg_fit else 20, # maximum value of parameter
delta=0.05, # initial step used by fitting engine
unit = u.Hz
)
Define a point source at the known location with a Band function spectrum and add it to the model. The initial values of the Band function parameters are set to the true values used to simulate the source
[9]:
l = 184.56
b = -5.78
alpha = -1.99
beta = -2.32
E0 = 531. * (alpha - beta) * u.keV
xp = E0 * (alpha + 2) / (alpha - beta)
piv = 500. * u.keV
K = 3.07e-5 / u.cm / u.cm / u.s / u.keV
spectrum = Band()
spectrum.alpha.min_value = -2.14
spectrum.alpha.max_value = 3.0
spectrum.beta.min_value = -5.0
spectrum.beta.max_value = -2.15
spectrum.xp.min_value = 1.0
spectrum.alpha.delta = 0.01
spectrum.beta.delta = 0.01
spectrum.alpha.value = alpha
spectrum.beta.value = beta
spectrum.xp.value = xp.value
spectrum.K.value = K.value
spectrum.piv.value = piv.value
spectrum.xp.unit = xp.unit
spectrum.K.unit = K.unit
spectrum.piv.unit = piv.unit
source = PointSource("source", # Name of source (arbitrary, but needs to be unique)
l = l, # Longitude (deg)
b = b, # Latitude (deg)
spectral_shape = spectrum) # Spectral model
# Optional: free the position parameters
#source.position.l.free = True
#source.position.b.free = True
model = Model(source) # Model with single source. If we had multiple sources, we would do Model(source1, source2, ...)
15:28:37 WARNING The current value of the parameter beta (-2.0) was above the new maximum -2.15. parameter.py:810
Gather all plugins and combine with the model in a JointLikelihood object, then perform maximum likelihood fit
[10]:
#[magic commented out by run_tutorials.py]%%time
plugins = DataList(cosi) # If we had multiple instruments, we would do e.g. DataList(cosi, lat, hawc, ...)
like = JointLikelihood(model, plugins, verbose = False)
_ = like.fit()
15:28:37 INFO set the minimizer to minuit joint_likelihood.py:994
WARNING IntegrationWarning: The occurrence of roundoff error is detected, which prevents
the requested tolerance from being achieved. The error may be
underestimated.
15:28:46 WARNING The current value of the parameter beta (-2.0) was above the new maximum -2.15. parameter.py:810
Best fit values:
| result | unit | |
|---|---|---|
| parameter | ||
| source.spectrum.main.Band.K | (5.41 +/- 0.11) x 10^-5 | 1 / (keV s cm2) |
| source.spectrum.main.Band.alpha | -1.838 +/- 0.005 | |
| source.spectrum.main.Band.xp | (1.78 +/- 0.09) x 10 | keV |
| source.spectrum.main.Band.beta | -2.2211 +/- 0.0034 | |
| total_bkg | (2.22876 +/- 0.00024) x 10 | Hz |
Correlation matrix:
| 1.00 | 0.60 | -0.23 | -0.06 | -0.03 |
| 0.60 | 1.00 | 0.59 | 0.16 | -0.01 |
| -0.23 | 0.59 | 1.00 | -0.06 | -0.02 |
| -0.06 | 0.16 | -0.06 | 1.00 | -0.19 |
| -0.03 | -0.01 | -0.02 | -0.19 | 1.00 |
Values of -log(likelihood) at the minimum:
| -log(likelihood) | |
|---|---|
| cosi | -1169452582.9882479 |
| total | -1169452582.9882479 |
Values of statistical measures:
| statistical measures | |
|---|---|
| AIC | -2338905155.9762354 |
| BIC | -2338905104.2386346 |
Error propagation and plotting
Define Band function spectrum injected into MEGAlib
[11]:
alpha_inj = -1.99
beta_inj = -2.32
E0_inj = 531. * (alpha_inj - beta_inj) * u.keV
xp_inj = E0_inj * (alpha_inj + 2) / (alpha_inj - beta_inj)
piv_inj = 100. * u.keV
K_inj = 7.56e-4 / u.cm / u.cm / u.s / u.keV
spectrum_inj = Band()
spectrum_inj.alpha.min_value = -2.14
spectrum_inj.alpha.max_value = 3.0
spectrum_inj.beta.min_value = -5.0
spectrum_inj.beta.max_value = -2.15
spectrum_inj.xp.min_value = 1.0
spectrum_inj.alpha.value = alpha_inj
spectrum_inj.beta.value = beta_inj
spectrum_inj.xp.value = xp_inj.value
spectrum_inj.K.value = K_inj.value
spectrum_inj.piv.value = piv_inj.value
spectrum_inj.xp.unit = xp_inj.unit
spectrum_inj.K.unit = K_inj.unit
spectrum_inj.piv.unit = piv_inj.unit
WARNING The current value of the parameter beta (-2.0) was above the new maximum -2.15. parameter.py:810
The summary of the results above tell you the optimal values of the parameters, as well as the errors. Propogate the errors to the “evaluate_at” method of the spectrum
[12]:
results = like.results
print(results.display())
parameters = {par.name:results.get_variates(par.path)
for par in results.optimized_model["source"].parameters.values()
if par.free}
results_err = results.propagate(results.optimized_model["source"].spectrum.main.shape.evaluate_at, **parameters)
print(results.optimized_model["source"])
Best fit values:
| result | unit | |
|---|---|---|
| parameter | ||
| source.spectrum.main.Band.K | (5.41 +/- 0.11) x 10^-5 | 1 / (keV s cm2) |
| source.spectrum.main.Band.alpha | -1.838 +/- 0.005 | |
| source.spectrum.main.Band.xp | (1.78 +/- 0.09) x 10 | keV |
| source.spectrum.main.Band.beta | -2.2211 +/- 0.0034 | |
| total_bkg | (2.22876 +/- 0.00024) x 10 | Hz |
Correlation matrix:
| 1.00 | 0.60 | -0.23 | -0.06 | -0.03 |
| 0.60 | 1.00 | 0.59 | 0.16 | -0.01 |
| -0.23 | 0.59 | 1.00 | -0.06 | -0.02 |
| -0.06 | 0.16 | -0.06 | 1.00 | -0.19 |
| -0.03 | -0.01 | -0.02 | -0.19 | 1.00 |
Values of -log(likelihood) at the minimum:
| -log(likelihood) | |
|---|---|
| cosi | -1169452582.9882479 |
| total | -1169452582.9882479 |
Values of statistical measures:
| statistical measures | |
|---|---|
| AIC | -2338905155.9762354 |
| BIC | -2338905104.2386346 |
None
WARNING The current value of the parameter beta (-2.0) was above the new maximum -2.15. parameter.py:810
WARNING The current value of the parameter beta (-2.0) was above the new maximum -2.15. parameter.py:810
WARNING The current value of the parameter beta (-2.0) was above the new maximum -2.15. parameter.py:810
* source (point source):
* position:
* l:
* value: 184.56
* desc: Galactic longitude
* min_value: 0.0
* max_value: 360.0
* unit: deg
* is_normalization: false
* b:
* value: -5.78
* desc: Galactic latitude
* min_value: -90.0
* max_value: 90.0
* unit: deg
* is_normalization: false
* equinox: J2000
* spectrum:
* main:
* Band:
* K:
* value: 5.4051037971774306e-05
* desc: Differential flux at the pivot energy
* min_value: 1.0e-50
* max_value: null
* unit: keV-1 s-1 cm-2
* is_normalization: true
* alpha:
* value: -1.8382906397995242
* desc: low-energy photon index
* min_value: -2.14
* max_value: 3.0
* unit: ''
* is_normalization: false
* xp:
* value: 17.80704053659518
* desc: peak in the x * x * N (nuFnu if x is a energy)
* min_value: 1.0
* max_value: null
* unit: keV
* is_normalization: false
* beta:
* value: -2.221118197373546
* desc: high-energy photon index
* min_value: -5.0
* max_value: -2.15
* unit: ''
* is_normalization: false
* piv:
* value: 500.0
* desc: pivot energy
* min_value: null
* max_value: null
* unit: keV
* is_normalization: false
* polarization: {}
Evaluate the flux and errors at a range of energies for the fitted and injected spectra, and the simulated source flux
[13]:
energy = np.geomspace(100*u.keV,10*u.MeV).to_value(u.keV)
flux_lo = np.zeros_like(energy)
flux_median = np.zeros_like(energy)
flux_hi = np.zeros_like(energy)
flux_inj = np.zeros_like(energy)
for i, e in enumerate(energy):
flux = results_err(e)
flux_median[i] = flux.median
flux_lo[i], flux_hi[i] = flux.equal_tail_interval(cl=0.68)
flux_inj[i] = spectrum_inj.evaluate_at(e)
binned_energy_edges = crab.binned_data.axes['Em'].edges.value
binned_energy = np.array([])
bin_sizes = np.array([])
for i in range(len(binned_energy_edges)-1):
binned_energy = np.append(binned_energy, (binned_energy_edges[i+1] + binned_energy_edges[i]) / 2)
bin_sizes = np.append(bin_sizes, binned_energy_edges[i+1] - binned_energy_edges[i])
expectation = response.expectation()
Plot the fitted and injected spectra
[14]:
fig,ax = plt.subplots()
ax.plot(energy, energy*energy*flux_median, label = "Best fit")
ax.fill_between(energy, energy*energy*flux_lo, energy*energy*flux_hi, alpha = .5, label = "Best fit (errors)")
ax.plot(energy, energy*energy*flux_inj, color = 'black', ls = ":", label = "Injected")
ax.set_xscale("log")
ax.set_yscale("log")
ax.set_xlabel("Energy (keV)")
ax.set_ylabel(r"$E^2 \frac{dN}{dE}$ (keV cm$^{-2}$ s$^{-1}$)")
ax.set_ylim(.1,100)
_ = ax.legend()
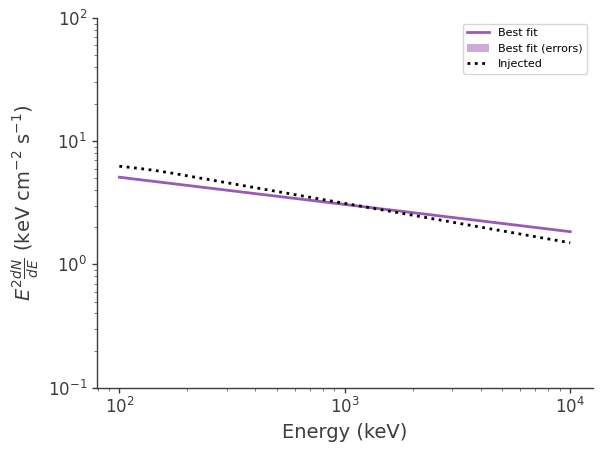
Plot the fitted spectrum convolved with the response, as well as the simulated source counts
[15]:
fig,ax = plt.subplots()
ax.stairs(expectation.project('Em').todense().contents, binned_energy_edges, color='purple', label = "Best fit convolved with response")
ax.errorbar(binned_energy, expectation.project('Em').todense().contents, yerr=np.sqrt(expectation.project('Em').todense().contents), color='purple', linewidth=0, elinewidth=1)
ax.stairs(crab.binned_data.project('Em').todense().contents, binned_energy_edges, color = 'black', ls = ":", label = "Source counts")
ax.errorbar(binned_energy, crab.binned_data.project('Em').todense().contents, yerr=np.sqrt(crab.binned_data.project('Em').todense().contents), color='black', linewidth=0, elinewidth=1)
ax.set_xscale("log")
ax.set_yscale("log")
ax.set_xlabel("Energy (keV)")
ax.set_ylabel("Counts")
_ = ax.legend()
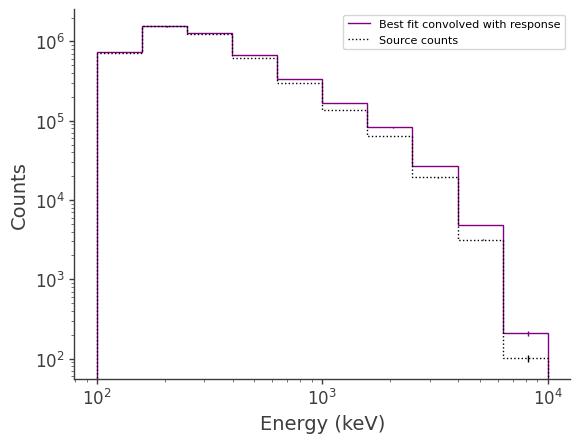
Plot the fitted spectrum convolved with the response plus the fitted background, as well as the simulated source+background counts
[16]:
expectation_bkg = bkg.expectation(copy = True)
fig,ax = plt.subplots()
ax.stairs(expectation.project('Em').todense().contents + expectation_bkg.project('Em').todense().contents, binned_energy_edges, color='purple', label = "Best fit convolved with response plus background")
ax.errorbar(binned_energy, expectation.project('Em').todense().contents+expectation_bkg.project('Em').todense().contents, yerr=np.sqrt(expectation.project('Em').todense().contents+expectation_bkg.project('Em').todense().contents), color='purple', linewidth=0, elinewidth=1)
ax.stairs(data.data.project('Em').todense().contents, binned_energy_edges, color = 'black', ls = ":", label = "Total counts")
ax.errorbar(binned_energy, data.data.project('Em').todense().contents, yerr=np.sqrt(data.data.project('Em').todense().contents), color='black', linewidth=0, elinewidth=1)
ax.set_xscale("log")
ax.set_yscale("log")
ax.set_xlabel("Energy (keV)")
ax.set_ylabel("Counts")
ax.legend()
plt.show()
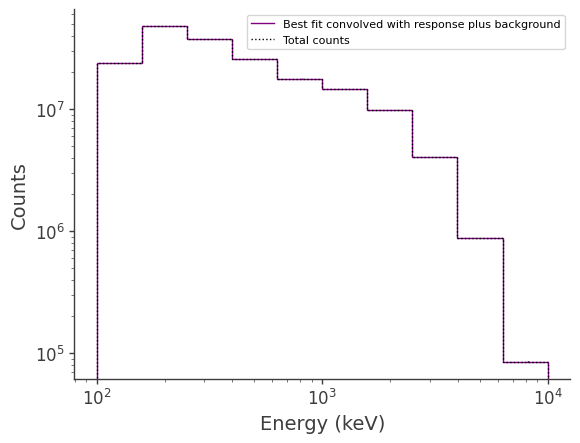
[ ]: